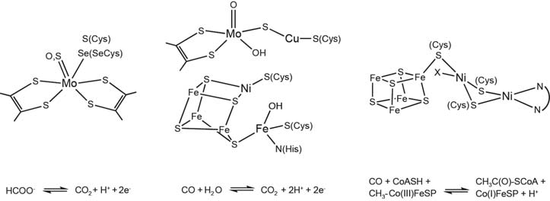
Carbon monoxide dehydrogenases
Carbon monoxide dehydrogenases (CODH) catalyse the oxidation of CO with H2O to yield CO2, two protons and two electrons. Two types of carbon monoxide dehydrogenases (CODHs) have been described: Ni- and Fe-dependent CODHs catalyse the oxidation of CO and the reduction of CO2 in a diverse set of microorganisms under anaerobic conditions. Mo- and Cu-dependent CODHs are able to oxidize CO and H2 and are found in bacteria dependent on aerobic respiration (see Figure). These enzymes are able to bind and to activate carbon oxides with high affinity and selectivity, and it is thus of interest to understand and to compare the different catalytic strategies for the two types of enzyme-bound heterometal centers.
We have developed an expression system for different CODHs from Carboxydothermus hydrogenoformans and established purification and crystallization conditions for the enzymes. Using protein X-ray crystallography several enzyme-adduct complexes have been analysed structurally and could be resolved at an atomic resolution (dmin < 1.2 Å). Due to the excellent resolution, this enzyme will be an ideal reference system for a thorough spectroscopic and theoretical analysis to elucidate the catalytic process.
Research goals
- Elucidating the factors (amino acids, ligands, oxidation state of metals) that control CO2 binding and reduction. EPR/ENDOR/ESEEM and IR spectroscopy will be specifically used to investigate the oxidation state of the Ni,Fe-cluster and the Ni-carbon bond of the CO2 bound state. The spectroscopic studies are complemented by quantum mechanical (QM) and hybrid QM/molecular mechanical (MM) methods.
- To determine the rate-limiting steps for the CO2-reduction, theoretical calculations will be combined with kinetic investigations employing rapid mixing techniques (stopped-flow and quenched-flow approaches) coupled to EPR and IR spectroscopic detection.
- As the interpretation of EPR spectroscopic data is currently hampered by the overlap with signals originating from the additional [4Fe-4S]-clusters in the CODHs, we will employ site directed mutagenesis to design CODH variants in which the signals of other paramagnetic centres are removed.
- Enzyme engineering will also be used to stabilize otherwise reactive states to facilitate their spectroscopic and structural characterisation and thus facilitate the theoretical description of the enzyme.
- We seek to develop a suitable expression system for CODH in E. coli to produce Cu-free apo-enzyme and to reconstitute the Cu center after purification of the enzyme. The ultimate goal is to obtain an expression system from which engineered mutants with substitutions in the active site can be prepared. Of particular interest is to determine the Mo-Cu site in different CODH variants and forms of the Cu-free enzyme, preferentially by X-ray crystallography. The determination of the Mo oxidation state will be supported by comparison with structural models synthesised in D2-1.
- Due to its oxygen-tolerance, the Mo-dependent enzyme has a potential as a CO sensor in biotechnological applications. Here, we will take the first steps in this direction by exploring the optimum conditions for immobilisation of the enzyme on conducting supports while preserving the native structure and efficient electronic coupling.
- Finally, it is planned to couple the Mo-containing CODH to a formate dehydrogenase (FDH) from E. coli by reversing the direction of the FDH reaction (see E2-2). Thus, formate will be produced from CO2 which in turn is the product of CODH. By coupling the oxygen-stable Mo-CODH with an oxygen-stable FDH, a CO-hydratase may be generated operating under aerobic conditions.